Jonathan Latham and Allison Wilson
Just before his appointment as head of the US National Institutes of Health (NIH), Francis Collins, the most prominent medical geneticist of our time, had his own genome scanned for disease susceptibility genes. He had decided, so he said, that the technology of personalised genomics was finally mature enough to yield meaningful results. Indeed, the outcome of his scan inspired The Language of Life, his recent book which urges every individual to do the same and secure their place on the personalised genomics bandwagon.
So, what knowledge did Collins’s scan produce? His results can be summarised very briefly. For North American males the probability of developing type 2 diabetes is 23%. Collins’s own risk was estimated at 29% and he highlighted this as the outstanding finding. For all other common diseases, however, including stroke, cancer, heart disease, and dementia, Collins’s likelihood of contracting them was average.
Predicting disease probability to within a percentage point might seem like a major scientific achievement. From the perspective of a professional geneticist, however, there is an obvious problem with these results. The hoped-for outcome is to detect genes that cause personal risk to deviate from the average. Otherwise, a genetic scan or even a whole genome sequence is showing nothing that wasn’t already known. The real story, therefore, of Collins’s personal genome scan is not its success, but rather its failure to reveal meaningful information about his long-term medical prospects. Moreover, Collins’s genome is unlikely to be an aberration. Contrary to expectations, the latest genetic research indicates that almost everyone’s genome will be similarly unrevealing.
We must assume that, as a geneticist as well as head of NIH, Francis Collins is more aware of this than anyone, but if so, he wrote The Language of Life not out of raw enthusiasm but because the genetics revolution (and not just personalised genomics) is in big trouble. He knows it is going to need all the boosters it can get.
What has changed scientifically in the last three years is the accumulating inability of a new whole-genome scanning technique (called Genome-Wide Association studies; GWAs) to find important genes for disease in human populations. In study after study, applying GWAs to every common (non-infectious) physical disease and mental disorder, the results have been remarkably consistent: only genes with very minor effects have been uncovered. In other words, the genetic variation confidently expected by medical geneticists to explain common diseases, cannot be found.
Read more
Sunday, 12 December 2010
Monday, 6 December 2010
Surprisingly sloppy yeast genes
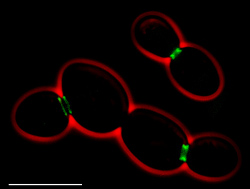
Contrary to popular belief, the gene expression of "housekeeping" proteins in yeast is not synchronized or even coordinated.
Instead, these essential genes -- which work together to build important cell complexes like ribosomes and proteasomes -- are turned on and off randomly, researchers report in today's online edition of Nature Structural and Molecular Biology.
"We all have our biases about how things work," said senior author Robert Singer of the Albert Einstein College of Medicine in New York. "Sometimes, we're just wrong."
The surprising finding may change the way scientists understand and assess some gene transcription networks.
"It's fantastic work," said Mads Kaern, Canada Research Chair in systems biology at the University of Ottawa, who was not involved in the research. "This challenges the idea that elaborate networks have evolved to regulate this class of genes. That is profound."
Multi-protein complexes -- like ribosomes, made up of 80 different proteins -- perform essential functions in cells. Scientists have long assumed the expression of the genes required to assemble such complexes is coordinated because often the resulting proteins have similar abundances in cells. Additionally, it made logical sense that widely varying levels of subunits would result in malformed complexes or a build-up of toxic levels of unused proteins.
But in 2008, Singer and colleagues studying the expression of genes in yeast noticed that many constitutive genes -- those transcribed throughout the cell cycle rather than on an as-needed basis -- had irregular, random expression patterns. The unexpected finding led them to predict that the expression of genes needed for multi-protein complexes might not be synchronized, either.
When the team measured the expression of several groups of genes -- those encoding subunits of a proteasome, a transcription factor, and RNA polymerase II -- they found that the mRNAs of each group's subunits were no more correlated than the mRNAs of genes that had no functional relationship to each other. Even two alleles of the same gene with identical promoters were not expressed equally.
"The genes are essentially clueless," said Singer. "They don't know what they're making or the actual destiny of protein. They're just there, cranking out proteins." In contrast, induced genes -- those triggered by stimuli, such as a nearby toxin or nutrient-rich media -- demonstrated highly coordinated expression.
Consequently, the researchers assume that the coordination of protein abundances for complexes happens after transcription. Because many proteins have longer half-lives than mRNA, for example, random fluctuations in mRNA levels may be inconsequential because proteins stick around for significantly longer. Or perhaps chaperone molecules impose checkpoints, stabilizing ribosomes or proteasomes to prevent dissociation until the next subunit arrives. Or both.
"We have this bias about cells being efficient, but the more we learn about them, the more inefficient we find out they are," said Singer. "But maybe that's the way biological systems have to work. If they had too many controls, there's a lot more opportunity for things to go wrong."
The un-coordinated expression of constitutive genes may have an important evolutionary function, suggested Kaern. "They may make cells more robust, less sensitive to mutations in upstream regulators," he said. If constitutive genes shared one upstream activator to turn them all on simultaneously, a single mutation in that activator could have catastrophic consequences for the cell, he speculated.
Overall, the research has important implications for systems biology and researchers studying gene networks of constitutive genes. "This opens up a conceptual door that was previously ignored," said Kaern, who suspects something similar may occur in mice and human cells. "But I think a lot of yeast biologists will not be surprised [this happens in yeast] because they know how sloppy yeast is," he added. "It has a tendency to survive whatever you throw at it."
Source: The Scientist
Thursday, 25 November 2010
Cancer’s conversions
A developmental transition may be a useful model for tumor progression.
Until recently, the universally accepted dogma in cancer research stated that replicating cells accumulate several rounds of mutations before becoming cancerous. According to that dogma, the mutations that result in metastatic spread throughout the body occur late in tumor progression. This idea has recently been challenged by the identification of cancer stem cells (CSCs), which provide a new explanation for both the initiation and propagation of tumorigenesis. Rather than following a linear process that starts with unchecked replication and ends with the loss of adhesion molecules that drives metastasis, CSCs can self-renew, proliferate, differentiate, and even revert back to a stem cell state, producing metastatic cells at unexpected stages of the disease.
With a new understanding that cancer progression does not necessarily follow a particular order, researchers have been looking for models to help explain how and when cancers become aggressive. While the idea of CSCs helps explain some observations that do not fit within the accepted dogma, researchers have proposed another idea based on a normal process in embryonic development called the epithelial-to-mesenchymal transition (EMT). Embryonic development requires epithelial cells to change gene expression patterns, lose their adhesion molecules, and become motile, mesenchymal-like cells that invade the extracellular matrix and later differentiate to form the interior tissues of the body such as skeletal muscles and the heart. A similar process, which includes the loss of adhesion molecules and the activation of common genes by Wnt/β-catenin and other signaling pathways, is also a prerequisite for malignant tumor development, particularly the metastatic process. Studying the basic biology of EMT, therefore, could shed light on the processes that cancer cells undergo as they develop.
Read more
Until recently, the universally accepted dogma in cancer research stated that replicating cells accumulate several rounds of mutations before becoming cancerous. According to that dogma, the mutations that result in metastatic spread throughout the body occur late in tumor progression. This idea has recently been challenged by the identification of cancer stem cells (CSCs), which provide a new explanation for both the initiation and propagation of tumorigenesis. Rather than following a linear process that starts with unchecked replication and ends with the loss of adhesion molecules that drives metastasis, CSCs can self-renew, proliferate, differentiate, and even revert back to a stem cell state, producing metastatic cells at unexpected stages of the disease.
With a new understanding that cancer progression does not necessarily follow a particular order, researchers have been looking for models to help explain how and when cancers become aggressive. While the idea of CSCs helps explain some observations that do not fit within the accepted dogma, researchers have proposed another idea based on a normal process in embryonic development called the epithelial-to-mesenchymal transition (EMT). Embryonic development requires epithelial cells to change gene expression patterns, lose their adhesion molecules, and become motile, mesenchymal-like cells that invade the extracellular matrix and later differentiate to form the interior tissues of the body such as skeletal muscles and the heart. A similar process, which includes the loss of adhesion molecules and the activation of common genes by Wnt/β-catenin and other signaling pathways, is also a prerequisite for malignant tumor development, particularly the metastatic process. Studying the basic biology of EMT, therefore, could shed light on the processes that cancer cells undergo as they develop.
Read more
Saturday, 20 November 2010
Induced global mutations
Imagine that some organisms find themselves in an environment in which they can no longer survive or reproduce. Their only hope of salvation is that a lucky mutation will crop up and enable them to deal with their adverse circumstances. If mutation rates are low, which they usually are, the chances that any will survive are slim. But if they have mechanisms that kick in in stressful conditions and increase the rate of mutation throughout the genome, things might be better.Many individuals will perish quickly (they get mutations that make matters worst), but the chances that one or two will have a liberating mutation are enhanced.
(...)
The mutation rates in bacteria are enhanced when they encounter an environment that is so hostile that they completely stop growing and reproducing. In such conditions, a spate of new mutations is generated throughout the genome. Every little single mutation is random, in the sense that it is not function-specific, but the general genomic response - the increased mutation rate - may be adaptive.
(...)
Not everyone accepts that stress-induced mutation is an evolved adaptation, however. Some people argue that the spate of mutations that occur in adverse conditions is simply a by-product of stress-induced failure. When cells are stressed, especially when starved, one of the things that may happen is that they are no longer able to produce the proteins needed for DNA maintenance and repair. It may even be that starved cells are obliged to turn off their DNA-caretaker genes to save energy. If so, faults will occur and remain uncorrected. In other words, there will be a lot of mutations. In this case, the generation of mutations is just a pathological symptom of the problems cells are experiencing, not an evolved adaptive response to adverse conditions.
From: Eva Jablonka & Marion Lamb - Evolution in four dimensions
(...)
The mutation rates in bacteria are enhanced when they encounter an environment that is so hostile that they completely stop growing and reproducing. In such conditions, a spate of new mutations is generated throughout the genome. Every little single mutation is random, in the sense that it is not function-specific, but the general genomic response - the increased mutation rate - may be adaptive.
(...)
Not everyone accepts that stress-induced mutation is an evolved adaptation, however. Some people argue that the spate of mutations that occur in adverse conditions is simply a by-product of stress-induced failure. When cells are stressed, especially when starved, one of the things that may happen is that they are no longer able to produce the proteins needed for DNA maintenance and repair. It may even be that starved cells are obliged to turn off their DNA-caretaker genes to save energy. If so, faults will occur and remain uncorrected. In other words, there will be a lot of mutations. In this case, the generation of mutations is just a pathological symptom of the problems cells are experiencing, not an evolved adaptive response to adverse conditions.
From: Eva Jablonka & Marion Lamb - Evolution in four dimensions
Wednesday, 17 November 2010
Molecular animations
Suggested by Glen Oomen (Medical Illustrator) & Serena Jennings (my colleague):
Sunday, 14 November 2010
Population thinking
Generalizations in biology are almost invariably of a probabilistic nature. As one wit has formulated it, there is only one universal law in biology: "All biological laws have exceptions". This probabilistic conceptualization contrasts strikingly with the view during the early period of the scientific revolution that causation in nature is regulated by laws that can be stated in mathematical terms. Actually, this idea occurred apparently first to Pythagoras. It has remained a dominant idea, particularly in the physical sciences, up to the present day. (...) With Plato, it gave rise to the essentialism. For him, the variable world of phenomena was nothing but the reflection of a limited number of fixed and unchanging forms, the essences. These essences are what is real and important in this world. Constancy and discontinuity are the points of special emphasis for the essentialists. Variation is attributed to the imperfect manifestation of the underlying essences.
(...)
Darwin, one of the first thinkers to reject the essentialism (at least in part), was not at all understood by the contemporary philosophers (all of whom were essentialists), and his concept of evolution through natural selection was therefore found unacceptable. Genuine change, according to essentialism, is possible only through the saltatation origin of new essences. Because evolution as explained by Darwin, is by necessity gradual, it is quite incompatible with essentialism.
Western thinking for more than two thousand years after Plato was dominated by essentialism. It was not until the nineteenth century that a new and different way of thinking about nature began to spread, so-called population thinking. What is population thinking and how does it differ from essentialism? Population thinkers stress the uniqueness of everything in the organic world. What is important for them is the individual, not the type. They emphasize that every individual in sexually reproducing species is uniquely different from all others, with much individuality even existing in uniparentally reproducing ones. There is no "typical" individual, and mean values are abstractions. Much of what in the past has been designated in biology as "classes" are populations consisting of unique individuals.
From Ernst Mayr - The Growth of Biological Thought
(...)
Darwin, one of the first thinkers to reject the essentialism (at least in part), was not at all understood by the contemporary philosophers (all of whom were essentialists), and his concept of evolution through natural selection was therefore found unacceptable. Genuine change, according to essentialism, is possible only through the saltatation origin of new essences. Because evolution as explained by Darwin, is by necessity gradual, it is quite incompatible with essentialism.
Western thinking for more than two thousand years after Plato was dominated by essentialism. It was not until the nineteenth century that a new and different way of thinking about nature began to spread, so-called population thinking. What is population thinking and how does it differ from essentialism? Population thinkers stress the uniqueness of everything in the organic world. What is important for them is the individual, not the type. They emphasize that every individual in sexually reproducing species is uniquely different from all others, with much individuality even existing in uniparentally reproducing ones. There is no "typical" individual, and mean values are abstractions. Much of what in the past has been designated in biology as "classes" are populations consisting of unique individuals.
From Ernst Mayr - The Growth of Biological Thought
Tuesday, 2 November 2010
Strong immunity = low fertility
A study in wild sheep may help explain why natural selection has not eradicated weak or self-destructive immune systems
A fluctuating trade-off between reproduction and survival in a feral population of Soay sheep may resolve the age-old question of why natural selection has failed to eradicate genes for both infection-prone and self-assailing immune systems.
The potential answer, published in Science this week (29th October), comes from the nascent field of ecoimmunology, which examines how different levels of antibodies in the blood of wild animals can influence their ability to survive and produce young.
Specifically, the authors found that, among a population of isolated, wild sheep, individuals with higher levels of antibodies associated with autoimmunity in other species were more likely to survive harsh weather conditions, but also reproduced less. Consequently, the benefits of high immunity, such as quick and efficient riddance of infection, may come with a cost -- less energy for reproduction.
"This paper reveals that more [antibodies] might not always be better, and that to understand the evolution of immune systems, it will be critical to study them in free-living, outbred organisms," Lynn Martin, an ecoimmunologist at the University of South Florida, who was not affiliated with the study, said in an email to The Scientist.
Researchers have found that feral rodents can hold comparatively high concentrations of antibodies in their blood, but, mysteriously, autoimmunity diseases such as type 1 diabetes and lupus are only seen in humans and lab, domestic, and captive mammals, said Andrea Graham, an evolutionary biologist at Princeton University and first author on the study.
There was "this nagging question of whether autoimmunity was some weird freak of captivity," added Andrew Read, evolutionary biologist at Pennsylvania State University, who did not contribute to the study.
Using blood samples collected every August for 11 years from the Soay sheep population on Hirta, an island in the St. Kilda archipelago of Scotland, Graham and colleagues from the University of Edinburgh in Scotland measured the concentration of antinuclear antibodies (ANAs), or autoimmune antibodies that attack the contents of the cell's nucleus as if it were foreign material. They then compared these levels to other variables of fitness such as survival and reproduction.
Researchers found that adult females with higher levels of ANAs lived longer by surviving more bitterly cold, parasite-infested winters. However, these same females were less likely to have babies the following spring. The correlation was only present during particularly harsh winters, however, when sometimes nearly 50 percent of the population died, suggesting heterogeneity in immune response is produced by natural selection acting on an ever-changing environment.
Ewes with high levels of ANAs also produced young with higher chances of survival through the next winter than young born to mothers with weak immune systems, suggesting a genetic basis for the varied immune responses in the sheep population.
"Immune response is only one part of the fitness component," said Peter Hudson, a disease ecologist at Pennsylvania State University, who was not affiliated with the study. "When [an individual] is not being exposed to pathogens, then high immune response could be too costly."
According to the results, when parasite prevalence is low and food is abundant, individuals with low immune responses will have the highest fitness because they will have the energy to produce the most young in the shortest period of time. But when the threat of infection is high and the winters are brutally frigid, individuals with high immune responses will survive and live on to have more children, while others die off. Thus, these trade-offs exhibited by the Soay sheep can account for their heterogeneity in immune response.
Read and Martin agreed that the next step is to experimentally manipulate antibody response in large populations to discover whether a causal, rather than correlational, relationship is present in this survival-reproduction trade-off.
The field of ecoimmunology "has been a controversial field because it's really hard to decide what to measure without a history [of the population]," noted Read.
This study demonstrates its potential benefits, however, Graham argued. "I firmly believe that we wouldn't have been able to find out such cool things about the immune system without this long study on the Soay sheep," she said. "Now that we understand all the nuts and bolts of the immune system [from traditional immunology], we can go on to try to understand it in the real world, because that's what really matters."
A. Graham et al., "Fitness Correlates of Heritable Variation in Antibody Responsiveness in a Wild Mammal," Science, 330:662-65, 2010.
Source: The Scientist
A fluctuating trade-off between reproduction and survival in a feral population of Soay sheep may resolve the age-old question of why natural selection has failed to eradicate genes for both infection-prone and self-assailing immune systems.
The potential answer, published in Science this week (29th October), comes from the nascent field of ecoimmunology, which examines how different levels of antibodies in the blood of wild animals can influence their ability to survive and produce young.
Specifically, the authors found that, among a population of isolated, wild sheep, individuals with higher levels of antibodies associated with autoimmunity in other species were more likely to survive harsh weather conditions, but also reproduced less. Consequently, the benefits of high immunity, such as quick and efficient riddance of infection, may come with a cost -- less energy for reproduction.
"This paper reveals that more [antibodies] might not always be better, and that to understand the evolution of immune systems, it will be critical to study them in free-living, outbred organisms," Lynn Martin, an ecoimmunologist at the University of South Florida, who was not affiliated with the study, said in an email to The Scientist.
Researchers have found that feral rodents can hold comparatively high concentrations of antibodies in their blood, but, mysteriously, autoimmunity diseases such as type 1 diabetes and lupus are only seen in humans and lab, domestic, and captive mammals, said Andrea Graham, an evolutionary biologist at Princeton University and first author on the study.
There was "this nagging question of whether autoimmunity was some weird freak of captivity," added Andrew Read, evolutionary biologist at Pennsylvania State University, who did not contribute to the study.
Using blood samples collected every August for 11 years from the Soay sheep population on Hirta, an island in the St. Kilda archipelago of Scotland, Graham and colleagues from the University of Edinburgh in Scotland measured the concentration of antinuclear antibodies (ANAs), or autoimmune antibodies that attack the contents of the cell's nucleus as if it were foreign material. They then compared these levels to other variables of fitness such as survival and reproduction.
Researchers found that adult females with higher levels of ANAs lived longer by surviving more bitterly cold, parasite-infested winters. However, these same females were less likely to have babies the following spring. The correlation was only present during particularly harsh winters, however, when sometimes nearly 50 percent of the population died, suggesting heterogeneity in immune response is produced by natural selection acting on an ever-changing environment.
Ewes with high levels of ANAs also produced young with higher chances of survival through the next winter than young born to mothers with weak immune systems, suggesting a genetic basis for the varied immune responses in the sheep population.
"Immune response is only one part of the fitness component," said Peter Hudson, a disease ecologist at Pennsylvania State University, who was not affiliated with the study. "When [an individual] is not being exposed to pathogens, then high immune response could be too costly."
According to the results, when parasite prevalence is low and food is abundant, individuals with low immune responses will have the highest fitness because they will have the energy to produce the most young in the shortest period of time. But when the threat of infection is high and the winters are brutally frigid, individuals with high immune responses will survive and live on to have more children, while others die off. Thus, these trade-offs exhibited by the Soay sheep can account for their heterogeneity in immune response.
Read and Martin agreed that the next step is to experimentally manipulate antibody response in large populations to discover whether a causal, rather than correlational, relationship is present in this survival-reproduction trade-off.
The field of ecoimmunology "has been a controversial field because it's really hard to decide what to measure without a history [of the population]," noted Read.
This study demonstrates its potential benefits, however, Graham argued. "I firmly believe that we wouldn't have been able to find out such cool things about the immune system without this long study on the Soay sheep," she said. "Now that we understand all the nuts and bolts of the immune system [from traditional immunology], we can go on to try to understand it in the real world, because that's what really matters."
A. Graham et al., "Fitness Correlates of Heritable Variation in Antibody Responsiveness in a Wild Mammal," Science, 330:662-65, 2010.
Source: The Scientist
Top 7 genetics papers
A snapshot of the highest-ranked articles in genetics and related areas in the past 30 days
1. Mapping transcriptomes
While mapping every transcriptional start site and operon of Helicobacter pylori at single-nucleotide resolution, the authors identify novel small RNAs, reveal the widespread nature of antisense transcription, and unveil a new technique to investigate the genomic complexities of other important pathogens, such as Salmonella and Mycobacterium tuberculosis.
2. Epigenetics in mind
The body's tendency to silence the expression of one parental allele in favor of the other -- a phenomenon known as genomic imprinting -- is much more widespread in the brain than scientists have believed, according to a new genome-wide study in mice. Surprisingly, more than 1300 genes expressed in the mouse brain appear to exhibit "parent-of-origin" epigenetic effects.
3. Translation goes local
Protein synthesis is a complicated game, but for the first time researchers have shown direct interaction between a transmembrane receptor, called DCC, and the translational machinery in rodent neurons, a step that likely facilitates localized protein production.
4. No RNA "dark matter"?
Most of the DNA that's transcribed into RNA in fact codes for proteins, a finding that disputes previous studies that suggested that the majority of mammalian transcripts are non-coding "dark matter."
5. Super E. Coli
The mother cell of E. coli maintains a constant growth rate throughout its replicative life (hundreds of cell divisions), despite accumulating damage and an increased probability of death, suggesting that growth and aging are decoupled, unlike all other studied aging models.
6. How autophagosomes form
Under conditions of starvation, autophagosomes form to resupply the cell by bringing nutrients from the cytosol or other organelles to the lysosomes, ensuring the cell's survival. New findings reveal an essential ingredient to this mysterious process: the outer membrane of mitochondria.
7. New tumor targets?
A scan of 1800 megabases of DNA from 441 tumors reveals more than 2500 somatic mutations, providing the mutation "spectra" for cancers, including protein kinases and G-protein-coupled receptors, some of which may serve as druggable targets.
Source: The scientist
1. Mapping transcriptomes
While mapping every transcriptional start site and operon of Helicobacter pylori at single-nucleotide resolution, the authors identify novel small RNAs, reveal the widespread nature of antisense transcription, and unveil a new technique to investigate the genomic complexities of other important pathogens, such as Salmonella and Mycobacterium tuberculosis.
2. Epigenetics in mind
The body's tendency to silence the expression of one parental allele in favor of the other -- a phenomenon known as genomic imprinting -- is much more widespread in the brain than scientists have believed, according to a new genome-wide study in mice. Surprisingly, more than 1300 genes expressed in the mouse brain appear to exhibit "parent-of-origin" epigenetic effects.
3. Translation goes local
Protein synthesis is a complicated game, but for the first time researchers have shown direct interaction between a transmembrane receptor, called DCC, and the translational machinery in rodent neurons, a step that likely facilitates localized protein production.
4. No RNA "dark matter"?
Most of the DNA that's transcribed into RNA in fact codes for proteins, a finding that disputes previous studies that suggested that the majority of mammalian transcripts are non-coding "dark matter."
5. Super E. Coli
The mother cell of E. coli maintains a constant growth rate throughout its replicative life (hundreds of cell divisions), despite accumulating damage and an increased probability of death, suggesting that growth and aging are decoupled, unlike all other studied aging models.
6. How autophagosomes form
Under conditions of starvation, autophagosomes form to resupply the cell by bringing nutrients from the cytosol or other organelles to the lysosomes, ensuring the cell's survival. New findings reveal an essential ingredient to this mysterious process: the outer membrane of mitochondria.
7. New tumor targets?
A scan of 1800 megabases of DNA from 441 tumors reveals more than 2500 somatic mutations, providing the mutation "spectra" for cancers, including protein kinases and G-protein-coupled receptors, some of which may serve as druggable targets.
Source: The scientist
What is systems biology?
For the fun of it, here are a few examples of definitions:
To understand complex biological systems requires the integration of experimental and computational research -- in other words a systems biology approach. (Kitano, 2002)
Systems biology studies biological systems by systematically perturbing them (biologically, genetically, or chemically); monitoring the gene, protein, and informational pathway responses; integrating these data; and ultimately, formulating mathematical models that describe the structure of the system and its response to individual perturbations. (Ideker et al, 2001)
[...]the objective of systems biology [can be] defined as the understanding of network behavior, and in particular their dynamic aspects, which requires the utilization of methematical modeling tightly linked to experiment. (Cassman, 2005)
By discovering how function arises in dynamic interactions, systems biology addresses the missing links between molecules and physiology. Top-down systems biology identifies molecular interaction networks on the basis of correlated molecular behavior observed in genome-wide "omics" studies. Bottom-up systems biology examines the mechanisms through which functional properties arise in the interactions of known components. (Bruggeman and Westerhoff, 2007)
Why is it so difficult to come up with a concise definition of systems biology? One of the reasons might be that every definition has to respect a delicate balance between "the yin and the yang" of the discipline: the integration of experimental and computational approaches; the balance between genome-wide systematical approaches and smaller-scale quantitative studies; top-down versus bottom-up strategies to solve systems architecture and functional properties. But despite the diversity in opinions and views, there might be two main aspects that are conserved across these definitions: a) a system-level approach attempts to consider all the components of a system; b) the properties and interactions of the components are linked with functions performed by the intact system via a computational model. This may in fact reveal another source of difficulty when trying to define systems biology, which is to find a general and objective definition of "biological function" (or Lander's "goal of the system", see our brief post Teleology and Systems Biology). Feel free to comment and suggest on this...
From http://blog-msb.embo.org/blog/2007/07/
Posted originally by Thomas on July 23, 2007
To understand complex biological systems requires the integration of experimental and computational research -- in other words a systems biology approach. (Kitano, 2002)
Systems biology studies biological systems by systematically perturbing them (biologically, genetically, or chemically); monitoring the gene, protein, and informational pathway responses; integrating these data; and ultimately, formulating mathematical models that describe the structure of the system and its response to individual perturbations. (Ideker et al, 2001)
[...]the objective of systems biology [can be] defined as the understanding of network behavior, and in particular their dynamic aspects, which requires the utilization of methematical modeling tightly linked to experiment. (Cassman, 2005)
By discovering how function arises in dynamic interactions, systems biology addresses the missing links between molecules and physiology. Top-down systems biology identifies molecular interaction networks on the basis of correlated molecular behavior observed in genome-wide "omics" studies. Bottom-up systems biology examines the mechanisms through which functional properties arise in the interactions of known components. (Bruggeman and Westerhoff, 2007)
Why is it so difficult to come up with a concise definition of systems biology? One of the reasons might be that every definition has to respect a delicate balance between "the yin and the yang" of the discipline: the integration of experimental and computational approaches; the balance between genome-wide systematical approaches and smaller-scale quantitative studies; top-down versus bottom-up strategies to solve systems architecture and functional properties. But despite the diversity in opinions and views, there might be two main aspects that are conserved across these definitions: a) a system-level approach attempts to consider all the components of a system; b) the properties and interactions of the components are linked with functions performed by the intact system via a computational model. This may in fact reveal another source of difficulty when trying to define systems biology, which is to find a general and objective definition of "biological function" (or Lander's "goal of the system", see our brief post Teleology and Systems Biology). Feel free to comment and suggest on this...
From http://blog-msb.embo.org/blog/2007/07/
Posted originally by Thomas on July 23, 2007
Thursday, 28 October 2010
Fine-grained exploration
Many, if not all, complex systems in biology have a fine-grained architecture, in that they consist of large numbers of relatively simple elements that work together in a highly parallel fashion.
Several possible advantages are conferred by this type of architecture, including robustness, efficiency, and evolvability. One additional major advantage is that a fine-grained parallel system is able to carry out what Douglas Hofstadter has called a "parallel terraced scan". This refers to a simultaneous exploration of many possibilities and pathways, in which the resources given to each exploration at a given time depend on the perceived success of that exploration at that time. The search is parallel in that many different possibilities are explored simultaneously, but is "terraced" in that not all possibilities are explored at the same speeds or to the same depth. Information is used as it is gained to continually reasses what is important to explore.
In cellular metabolism such fine-grained explorations are carried out by metabolic pathways, each focused on carrying out a particular task. A pathway can be speeded up or slowed down via feedback from its own results or from other pathways. The feedback itself is in the form of time-varying concentrations of molecules, so the relative speeds of different pathways can continually adapt to the moment-to-moment needs of the cell.
Melanie Mitchell - Complexity: a guided tour.
Several possible advantages are conferred by this type of architecture, including robustness, efficiency, and evolvability. One additional major advantage is that a fine-grained parallel system is able to carry out what Douglas Hofstadter has called a "parallel terraced scan". This refers to a simultaneous exploration of many possibilities and pathways, in which the resources given to each exploration at a given time depend on the perceived success of that exploration at that time. The search is parallel in that many different possibilities are explored simultaneously, but is "terraced" in that not all possibilities are explored at the same speeds or to the same depth. Information is used as it is gained to continually reasses what is important to explore.
In cellular metabolism such fine-grained explorations are carried out by metabolic pathways, each focused on carrying out a particular task. A pathway can be speeded up or slowed down via feedback from its own results or from other pathways. The feedback itself is in the form of time-varying concentrations of molecules, so the relative speeds of different pathways can continually adapt to the moment-to-moment needs of the cell.
Melanie Mitchell - Complexity: a guided tour.
Friday, 22 October 2010
Cheating yeast help group.
New results show that yeast populations grow better when a few individuals cheat the system
From: http://www.the-scientist.com
Yeast colonies with mooches, thieves and cheats actually grow faster and larger than colonies without these freeloading individuals, according to a study published 15th September in PLoS Biology, challenging the widely held belief that cheaters bring only bad news to cooperating populations.
Researchers found that when some yeast cheat their neighbors out of glucose, the entire population grows faster. Image: Eric Miller, Max Planck Institute of Evolutionary Biology "This is a most surprising result," said Laurence Hurst of the University of Bath in the UK, who coauthored the study. "The theory of cooperation was one of the best worked theories in all of evolution. Everyone assumed that it had to be the case that the world is better off when everyone cooperates."
The results may explain why yeast populations tolerate the presence of cheaters, added Michael Travisano, a biologist at the University of Minnesota, who was not involved in the research -- "because a mixed strategy is to everyone's benefit."
Most yeast secrete invertase, which hydrolyzes sucrose into fructose and glucose, their preferred food. However, some yeast are known to cheat the system. Cheater yeast don't secrete invertase and therefore don't contribute to the glucose production, yet they still eat the glucose that is generated by the rest of the population.
According to the theory of cooperation, which states that organisms are better off when everyone cooperates, yeast populations should be best off when all the yeast produce invertase. This would maximize the availability of glucose, which should enable more yeast growth. But when Hurst and his colleagues grew yeast populations with both producers and non-producers of invertase, this is not what they saw. Instead, the yeast grew the fastest and saw the highest population numbers when a proportion of the population was cheating.
One reason populations with cheaters grew better has to do with the yeast's inability to efficiently use abundant resources, Hurst said. "If you can hop down to your local McDonald's for a Big Mac, and it's very easy and very cheap, then you don't mind if you eat half of it and throw the rest away," Hurst said. "If you were starving in Africa, you wouldn't even imagine doing that." With the cheater yeast using up some of the available glucose, the cooperators are able to use the remaining resources much more efficiently, he said, allowing the population to grow larger and more quickly.
The team also modeled the experimental results in an effort to see whether their findings were specific to yeast growing on a petri dish, or whether they might apply to other organisms as well. The results showed that their experimental outcomes could be generalized -- cheats would benefit a population whenever certain criteria were met. These results imply that cooperation isn't always the most beneficial path for a population, Hurst said. Instead, the benefits of cooperation depend on the characteristics of the population itself. Under certain conditions, some amount of cheating is likely beneficial.
But the story is not a simple one, said Jeff Gore, a biophysicist at the Massachusetts Institute of Technology, who did not participate in the research. For example, "if the cheaters and cooperators are growing at different rates, the ratio of cooperators to cheaters won't be stable," Gore said. Thus, the population may be changing, and "you still have to ask what [it] is going to evolve to," and not just look at where it is now.
R.C. MacLean, et al., "A mixture of "cheats" and "cooperators" can enable maximal group benefit," PLoS Biol, 8(9): e1000486, 2010.
From: http://www.the-scientist.com
Yeast colonies with mooches, thieves and cheats actually grow faster and larger than colonies without these freeloading individuals, according to a study published 15th September in PLoS Biology, challenging the widely held belief that cheaters bring only bad news to cooperating populations.
Researchers found that when some yeast cheat their neighbors out of glucose, the entire population grows faster. Image: Eric Miller, Max Planck Institute of Evolutionary Biology "This is a most surprising result," said Laurence Hurst of the University of Bath in the UK, who coauthored the study. "The theory of cooperation was one of the best worked theories in all of evolution. Everyone assumed that it had to be the case that the world is better off when everyone cooperates."
The results may explain why yeast populations tolerate the presence of cheaters, added Michael Travisano, a biologist at the University of Minnesota, who was not involved in the research -- "because a mixed strategy is to everyone's benefit."
Most yeast secrete invertase, which hydrolyzes sucrose into fructose and glucose, their preferred food. However, some yeast are known to cheat the system. Cheater yeast don't secrete invertase and therefore don't contribute to the glucose production, yet they still eat the glucose that is generated by the rest of the population.
According to the theory of cooperation, which states that organisms are better off when everyone cooperates, yeast populations should be best off when all the yeast produce invertase. This would maximize the availability of glucose, which should enable more yeast growth. But when Hurst and his colleagues grew yeast populations with both producers and non-producers of invertase, this is not what they saw. Instead, the yeast grew the fastest and saw the highest population numbers when a proportion of the population was cheating.
One reason populations with cheaters grew better has to do with the yeast's inability to efficiently use abundant resources, Hurst said. "If you can hop down to your local McDonald's for a Big Mac, and it's very easy and very cheap, then you don't mind if you eat half of it and throw the rest away," Hurst said. "If you were starving in Africa, you wouldn't even imagine doing that." With the cheater yeast using up some of the available glucose, the cooperators are able to use the remaining resources much more efficiently, he said, allowing the population to grow larger and more quickly.
The team also modeled the experimental results in an effort to see whether their findings were specific to yeast growing on a petri dish, or whether they might apply to other organisms as well. The results showed that their experimental outcomes could be generalized -- cheats would benefit a population whenever certain criteria were met. These results imply that cooperation isn't always the most beneficial path for a population, Hurst said. Instead, the benefits of cooperation depend on the characteristics of the population itself. Under certain conditions, some amount of cheating is likely beneficial.
But the story is not a simple one, said Jeff Gore, a biophysicist at the Massachusetts Institute of Technology, who did not participate in the research. For example, "if the cheaters and cooperators are growing at different rates, the ratio of cooperators to cheaters won't be stable," Gore said. Thus, the population may be changing, and "you still have to ask what [it] is going to evolve to," and not just look at where it is now.
R.C. MacLean, et al., "A mixture of "cheats" and "cooperators" can enable maximal group benefit," PLoS Biol, 8(9): e1000486, 2010.
Saturday, 16 October 2010
Origins of organismal complexity
The vast majority of biologists engaged in evolutionary studies interpret virtually every aspect of biodiversity in adaptive terms. This narrow view of evolution has become untenable in light of recent observations from genomic sequencing and population-genetic theory. Numerous aspects of genomic architecture, gene structure, and developmental pathways are difficult to explain without invoking the nonadaptive forces of genetic drift and mutation. In addition, emergent biological features such as complexity, modularity, and evolvability, all of which are current targets of considerable speculation, may be nothing more than indirect by-products of processes operating at lower levels of organization. These issues are examined in the context of the view that the origins of many aspects of biological diversity, from gene-structural embellishments to novelties at the phenotypic level, have roots in nonadaptive processes, with the population-genetic environment imposing strong directionality on the paths that are open to evolutionary exploitation.
(...)
Although the basic theoretical foundation for understanding the mechanisms of evolution, the field of population genetics, has long been in place, the central significance of this framework is still occasionally questioned, as exemplified in this quote from Carroll, “Since the Modern Synthesis, most expositions of the evolutionary process have focused on microevolutionary mechanisms. Millions of biology students have been taught the view (from population genetics) that ‘evolution is change in gene frequencies.’ Isn't that an inspiring theme? This view forces the explanation toward mathematics and abstract descriptions of genes, and away from butterflies and zebras…. The evolution of form is the main drama of life's story, both as found in the fossil record and in the diversity of living species. So, let's teach that story. Instead of ‘change in gene frequencies,’ let's try ‘evolution of form is change in development’.” Even ignoring the fact that most species are unicellular and differentiated mainly by metabolic features, this statement illustrates two fundamental misunderstandings. Evolutionary biology is not a story-telling exercise, and the goal of population genetics is not to be inspiring, but to be explanatory. The roots of this contention are fourfold.
First, evolution is a population-genetic process governed by four fundamental forces. Darwin articulated one of those forces, the process of natural selection, for which an elaborate theory in terms of genotype frequencies now exists. The remaining three evolutionary forces are nonadaptive in the sense that they are not a function of the fitness properties of individuals: mutation is the ultimate source of variation on which natural selection acts, recombination assorts variation within and among chromosomes, and genetic drift ensures that gene frequencies will deviate a bit from generation to generation independent of other forces. Given the century of work devoted to the study of evolution, it is reasonable to conclude that these four broad classes encompass all of the fundamental forces of evolution. From Michael Lynch. PNAS May 15, 2007 vol. 104 no. Suppl 1 8597-8604.
(...)
Although the basic theoretical foundation for understanding the mechanisms of evolution, the field of population genetics, has long been in place, the central significance of this framework is still occasionally questioned, as exemplified in this quote from Carroll, “Since the Modern Synthesis, most expositions of the evolutionary process have focused on microevolutionary mechanisms. Millions of biology students have been taught the view (from population genetics) that ‘evolution is change in gene frequencies.’ Isn't that an inspiring theme? This view forces the explanation toward mathematics and abstract descriptions of genes, and away from butterflies and zebras…. The evolution of form is the main drama of life's story, both as found in the fossil record and in the diversity of living species. So, let's teach that story. Instead of ‘change in gene frequencies,’ let's try ‘evolution of form is change in development’.” Even ignoring the fact that most species are unicellular and differentiated mainly by metabolic features, this statement illustrates two fundamental misunderstandings. Evolutionary biology is not a story-telling exercise, and the goal of population genetics is not to be inspiring, but to be explanatory. The roots of this contention are fourfold.
First, evolution is a population-genetic process governed by four fundamental forces. Darwin articulated one of those forces, the process of natural selection, for which an elaborate theory in terms of genotype frequencies now exists. The remaining three evolutionary forces are nonadaptive in the sense that they are not a function of the fitness properties of individuals: mutation is the ultimate source of variation on which natural selection acts, recombination assorts variation within and among chromosomes, and genetic drift ensures that gene frequencies will deviate a bit from generation to generation independent of other forces. Given the century of work devoted to the study of evolution, it is reasonable to conclude that these four broad classes encompass all of the fundamental forces of evolution. From Michael Lynch. PNAS May 15, 2007 vol. 104 no. Suppl 1 8597-8604.
Friday, 15 October 2010
Evolution of genetic networks
Posted by Thomas in http://blog-msb.embo.org/blog/e/evolution_1/
A few days ago, an exciting review by Michael Lynch was published in Nature Reviews Genetics (The evolution of genetic networks by non-adaptive processes, Lynch 2007a ), a close follow-up of another review, published in PNAS a few months ago (The frailty of adaptive hypotheses for the origins of organismal complexity, Lynch 2007b). Michael Lynch has also written a book on the topic: The Origins of Genome Architecture (read a review)
The architecture of biological networks are often hypothesized as being "shaped" by adaptive evolution to confer global properties such as redundancy, robustness, modularity, complexity and evolvability. Lynch has some robust comments (others have some too, see Jonathan Eisen's "adaptationomics awards") on the “vast majority of biologists engaged in evolutionary studies [who] interpret virtually every aspect of biodiversity in adaptive terms” (Lynch 2007b). In contrast to what he perceives as a widespread belief, Lynch states clearly:
It is an open question as to whether pathway complexity is a necessary prerequisite for the evolution of complex phenotypes, or whether the genome architectures of multicellular species are simply more conducive to the passive emergence of network connections.(Lynch 2007a)
Beyond its somewhat controversial tone, Lynch's central lesson is the need to adopt a population genetics viewpoint (“nothing in evolution makes sense except in light of population genetics”) and he reminds us that, beside natural selection, three additional non-adaptive processes drive the evolution of living organisms: genetic drift, mutation and recombination. By analyzing the interplay between relative rates of loss and gain of regulatory sites (which depend both on mutation rate and mutational target size such as non-coding DNA), population size and recombination frequency, he demonstrates that purely non-adaptive forces can, in principle, determine the level of connectivity of regulatory networks--for example, determine the predominance of highly connected network motifs over linear pathways--without invoking any inherent advantages of the respective architectures on biological functions related for example to development or metabolism. It appears thus that, depending on the population genetics parameters, network structure can be profoundly "shaped" by the mere physical processes of mutation and recombination. At the very least, Lynch proposes that such models should be considered as "null hypothesis" when claiming that selection is engaged in a given aspect of organisms complexity.
In his review of Lynch's book, Massimo Pigliucci draws our attention to the fact that "the genome is only part of the story, arguably the simplest part to figure out", and that one of the greatest current challenges is to explain how phenotypes evolve. Lynch also recognizes that his models are simplified and do not, for example, consider kinetic or dynamical properties of biological networks. But here is a naive question: would it be possible to design an experimental strategy to test directly, in the lab, the evolution of simple (synthetic?) genetic circuits and observe the trends in connectivity under non-selective conditions or are the timescales involved too unrealistic?
A few days ago, an exciting review by Michael Lynch was published in Nature Reviews Genetics (The evolution of genetic networks by non-adaptive processes, Lynch 2007a ), a close follow-up of another review, published in PNAS a few months ago (The frailty of adaptive hypotheses for the origins of organismal complexity, Lynch 2007b). Michael Lynch has also written a book on the topic: The Origins of Genome Architecture (read a review)
The architecture of biological networks are often hypothesized as being "shaped" by adaptive evolution to confer global properties such as redundancy, robustness, modularity, complexity and evolvability. Lynch has some robust comments (others have some too, see Jonathan Eisen's "adaptationomics awards") on the “vast majority of biologists engaged in evolutionary studies [who] interpret virtually every aspect of biodiversity in adaptive terms” (Lynch 2007b). In contrast to what he perceives as a widespread belief, Lynch states clearly:
It is an open question as to whether pathway complexity is a necessary prerequisite for the evolution of complex phenotypes, or whether the genome architectures of multicellular species are simply more conducive to the passive emergence of network connections.(Lynch 2007a)
Beyond its somewhat controversial tone, Lynch's central lesson is the need to adopt a population genetics viewpoint (“nothing in evolution makes sense except in light of population genetics”) and he reminds us that, beside natural selection, three additional non-adaptive processes drive the evolution of living organisms: genetic drift, mutation and recombination. By analyzing the interplay between relative rates of loss and gain of regulatory sites (which depend both on mutation rate and mutational target size such as non-coding DNA), population size and recombination frequency, he demonstrates that purely non-adaptive forces can, in principle, determine the level of connectivity of regulatory networks--for example, determine the predominance of highly connected network motifs over linear pathways--without invoking any inherent advantages of the respective architectures on biological functions related for example to development or metabolism. It appears thus that, depending on the population genetics parameters, network structure can be profoundly "shaped" by the mere physical processes of mutation and recombination. At the very least, Lynch proposes that such models should be considered as "null hypothesis" when claiming that selection is engaged in a given aspect of organisms complexity.
In his review of Lynch's book, Massimo Pigliucci draws our attention to the fact that "the genome is only part of the story, arguably the simplest part to figure out", and that one of the greatest current challenges is to explain how phenotypes evolve. Lynch also recognizes that his models are simplified and do not, for example, consider kinetic or dynamical properties of biological networks. But here is a naive question: would it be possible to design an experimental strategy to test directly, in the lab, the evolution of simple (synthetic?) genetic circuits and observe the trends in connectivity under non-selective conditions or are the timescales involved too unrealistic?
Monday, 11 October 2010
Modularity and evolution
It may not be easy for one species to change into another. Imagine fish evolving to become dry land species. It is easy to see how problems arise as fins become legs and lungs are needed. Somehow, the evolutionary process got there. How? Almost certainly, the features of modularity and redundancy were critical. It is possible to slot pre-existing gene-protein networks into new control networks without upsetting them too much. That is modularity.
Suppose mutations occur in some mechanisms, and eventually they are selected for functions quite different from those they originally supported. How, then, to maintain the function that they originally served? What were back-up mechanisms now become primary. That is the value of redundancy. This is the basic explanation for how nature can modify its "aircraft design" while still ensuring that the aircraft continues to fly. By Denis Noble: The music of life - Biology beyond genes.
Suppose mutations occur in some mechanisms, and eventually they are selected for functions quite different from those they originally supported. How, then, to maintain the function that they originally served? What were back-up mechanisms now become primary. That is the value of redundancy. This is the basic explanation for how nature can modify its "aircraft design" while still ensuring that the aircraft continues to fly. By Denis Noble: The music of life - Biology beyond genes.
Fascinating correlations or elegant theories?
From: http://blog-msb.embo.org/blog/ Posted by Thomas on July 10, 2008
Chris Anderson, Editor-in-Chief of Wired , wrote a few weeks ago a provocative piece "The End of Theory: The Data Deluge Makes the Scientific Method Obsolete", arguing that in our Google-driven data-rich era ("The Petabyte Age") the good old "approach to science —hypothesize, model, test — is becoming obsolete", leaving place to a purely correlative vision of the world. There is a good dose of provocation in the essay and it was quite successful in spurring a flurry of skeptical reactions in the blogosphere, FriendFeed-land and lately in Edge's Reality Club.
I know that it is a bit late to write a post on this but this debate reminds me of the bottom-up vs top-down dialectic in (systems) biology. The tradition in molecular biology has been to focus on molecular mechanisms–a series of molecular events–that explain given biological functions. With detailed knowledge on the properties of an increasing number of components, bottom-up mechanistic descriptions–or models–can be constructed, which account for the experimental observations.
Of course, the purpose of models, at least for insightful ones, is more than merely providing mechanistic descriptions. As William Bialek writes, "Given a progressively more complete microscopic description of proteins and their interactions, how do we understand the emergence of function?" (Aguera y Arcas et al, 2003). There is therefore some subsequent subtle transition from description to insight, from model to theory, from detailed and specific to simple and general (watch Murray Gell-Mann's TEDTalk on "Beauty and truth in physics").
Theories are elegant.
On the other hand, high-throughput technologies (microarrays, proteomics, metabolomics, ultra high throughput sequencing, etc...) are indeed profoundly changing molecular biology and flooding the field with experimental data like never before. Currently, only part of this data can be explained within the context of mechanistic models. Still, and this is probably Chris Anderson's main point, it turns out that if the data is rich enough, one can exploit it by looking at the data globally, from the 'top', to reveal statistical patterns and correlations. Even if there is no mechanistic explanations (yet) for these correlations, they may reveal new worlds, novel structures and detect relationships between processes that were considered before as unlinked.
Correlations are fascinating.
Correlations resulting from data-driven analysis may well in turn stimulate new mechanistic investigations and hopefully new understanding. On Edge, Sean Carroll summarizes it all: "Sometimes it will be hard, or impossible, to discover simple models explaining huge collections of messy data taken from noisy, nonlinear phenomenon. But it doesn't mean we shouldn't try. Hypotheses aren't simply useful tools in some potentially-outmoded vision of science; they are the whole point. Theory is understanding, and understanding our world is what science is all about."
BUT, what is true for fundamental science is not obligatorily a rule for more applied fields, where the priority might less be on understanding than on acting. In particular, in medically related fields, top-down data-driven correlative approaches represent a pragmatic approach to obtain predictive models without waiting for still elusive fully mechanistic models that would encompass the entire complexity of human physiology (Nicholson, 2006).
As often in science, as in other human activities, different but complementary views are championed by people with different temperaments: there are those who like to build an edifice piece by piece and those who want to explore new territories. I think–I hope–that progresses in systems biology on both fronts, top-down and bottom-up, demonstrates that there is no need to turn this complementarity into an opposition.
Chris Anderson, Editor-in-Chief of Wired , wrote a few weeks ago a provocative piece "The End of Theory: The Data Deluge Makes the Scientific Method Obsolete", arguing that in our Google-driven data-rich era ("The Petabyte Age") the good old "approach to science —hypothesize, model, test — is becoming obsolete", leaving place to a purely correlative vision of the world. There is a good dose of provocation in the essay and it was quite successful in spurring a flurry of skeptical reactions in the blogosphere, FriendFeed-land and lately in Edge's Reality Club.
I know that it is a bit late to write a post on this but this debate reminds me of the bottom-up vs top-down dialectic in (systems) biology. The tradition in molecular biology has been to focus on molecular mechanisms–a series of molecular events–that explain given biological functions. With detailed knowledge on the properties of an increasing number of components, bottom-up mechanistic descriptions–or models–can be constructed, which account for the experimental observations.
Of course, the purpose of models, at least for insightful ones, is more than merely providing mechanistic descriptions. As William Bialek writes, "Given a progressively more complete microscopic description of proteins and their interactions, how do we understand the emergence of function?" (Aguera y Arcas et al, 2003). There is therefore some subsequent subtle transition from description to insight, from model to theory, from detailed and specific to simple and general (watch Murray Gell-Mann's TEDTalk on "Beauty and truth in physics").
Theories are elegant.
On the other hand, high-throughput technologies (microarrays, proteomics, metabolomics, ultra high throughput sequencing, etc...) are indeed profoundly changing molecular biology and flooding the field with experimental data like never before. Currently, only part of this data can be explained within the context of mechanistic models. Still, and this is probably Chris Anderson's main point, it turns out that if the data is rich enough, one can exploit it by looking at the data globally, from the 'top', to reveal statistical patterns and correlations. Even if there is no mechanistic explanations (yet) for these correlations, they may reveal new worlds, novel structures and detect relationships between processes that were considered before as unlinked.
Correlations are fascinating.
Correlations resulting from data-driven analysis may well in turn stimulate new mechanistic investigations and hopefully new understanding. On Edge, Sean Carroll summarizes it all: "Sometimes it will be hard, or impossible, to discover simple models explaining huge collections of messy data taken from noisy, nonlinear phenomenon. But it doesn't mean we shouldn't try. Hypotheses aren't simply useful tools in some potentially-outmoded vision of science; they are the whole point. Theory is understanding, and understanding our world is what science is all about."
BUT, what is true for fundamental science is not obligatorily a rule for more applied fields, where the priority might less be on understanding than on acting. In particular, in medically related fields, top-down data-driven correlative approaches represent a pragmatic approach to obtain predictive models without waiting for still elusive fully mechanistic models that would encompass the entire complexity of human physiology (Nicholson, 2006).
As often in science, as in other human activities, different but complementary views are championed by people with different temperaments: there are those who like to build an edifice piece by piece and those who want to explore new territories. I think–I hope–that progresses in systems biology on both fronts, top-down and bottom-up, demonstrates that there is no need to turn this complementarity into an opposition.
Genetics and the human nature
It's sometimes said that the genes determine the limits up to which, but not beyond which, a person's development may advance. This only confuses the issue. There is no way to predict all the phenotypes that a given genotype might yield in every one of the infinity of possible environments. Environments are infinitely diversified, and in the future there will exist environments that do not exist now. (...) Heredity cannot be called the "dice of destiny". Variations in body build, in physiology, and in mental traits are in part genetically conditioned, but this does not make education and social improvements any less well founded. What genetic conditioning does mean is that there is no single human nature, only human natures with different requirements for optimal growth and self-realization. The evidence of genetic conditioning of human traits, especially mental traits, must be examined with the greatest care. Theodosius Dobzhansky - Mankind evolving (1962).
Tuesday, 5 October 2010
Cis-regulatory evolution
Animal genomes are littered with conserved non-coding elements (CNEs) - most of which represent evolutionarily constrained cis-regulatory sequences - however, it is often not clear why these sequences are so exceptionally conserved, since anecdotal examples have shown that orthologous CNEs can have divergent functions in vivo. In an article recently published in Molecular Biology & Evolution, Ritter et al. compare the functional activities of 41 pairs of orthologous conserved non-coding elements (CNEs) from humans and zebrafish (2010). Interestingly, sequence similarity was found to be a poor predictor of which CNEs had conserved function. In contrast, the authors found that measuring transcription factor binding site change, instead of simple sequence divergence, improves their ability to predict functional conservation. While this set of tested CNEs remains relatively small, these results are encouraging because they suggest that as scientists move from phenomenological measures of CNE evolution to models based explicitly on binding site evolution, the patterns of cis-regulatory evolution observed within animal genomes should become far less mysterious.
From: http://blog-msb.embo.org/blog/
From: http://blog-msb.embo.org/blog/
Sunday, 19 September 2010
Living systems are not intentionally orchestrated
Living systems are simply not as they would be if all their components parts had been intentionally orchestrated. So when we talk of someone who "plays" the genome, we must recognise that no one set of molecules is given a position of privilege over another. Nature always use whatever comes to hand. That is also how it has evolved the pattern of regulation that we can to some extent imagine as the "organist". The components of a system may survive only because that higher-order system conforms to a particular, successful logic of survival of the organism, but that does not mean each component operates in an ideal way, or the best way in conformity with that logic. Indeed the system has to accommodate all sorts of lower-level quirks to be viable at all. Enrico Coen put it well. "Organisms", he said, "are not simply manufactured according to a set of instructions. There's no easy way to separate instructions from the process of carrying them out, to distinguish plan from execution".
From "The music of life - biology beyond genes" by Denis Noble
From "The music of life - biology beyond genes" by Denis Noble
Sunday, 22 August 2010
Defining and measuring complexity
What's complexity? For example, is the human genome more complex than the yeast genome (see my post on August 8th, 2010)? We intuitively answer this question with a big "OF COURSE". However, it has been surprisingly difficult to come up with an universally accepted definition of complexity. Although there is not yet a single science of complexity but rather several different sciences of complexity with different views about what complexity really means, the history of science shows us that the lack of an universally accepted definition of a central term in a new scientific field is more common than not. As an example, the modern genetics still does not have a good definition of gene at the molecular level.
The physicist Seth Lloyd proposed in 2001 three different dimensions along which to measure the complexity of a system:
1) How hard is it to describe?
2) How hard is it to create?
3) What is its degree of organization?
Another interesting proposed measure of complexity is the Shannon entropy, defined as the average information or "amount of surprise" a message source has for a receiver. Thus, using a classical example of genetics, we could say that the sequence CGTGGT has more entropy than the sequence AAAAAA and, therefore is more complex than the latter one. A completely random sequence has the maximum possible entropy. That means we could well make up an artificial genome by choosing a bunch of random As, Cs, Ts, and Gs. Using entropy as the measure of complexity, this random, almost certainly nonfunctional genome would be considered more complex than the human genome.
In conclusion: the most complex entities are not the most ordered or random ones but somewhere in between.
For further reading, see "Complexity: a guided tour" by M. Mitchell.
The physicist Seth Lloyd proposed in 2001 three different dimensions along which to measure the complexity of a system:
1) How hard is it to describe?
2) How hard is it to create?
3) What is its degree of organization?
Another interesting proposed measure of complexity is the Shannon entropy, defined as the average information or "amount of surprise" a message source has for a receiver. Thus, using a classical example of genetics, we could say that the sequence CGTGGT has more entropy than the sequence AAAAAA and, therefore is more complex than the latter one. A completely random sequence has the maximum possible entropy. That means we could well make up an artificial genome by choosing a bunch of random As, Cs, Ts, and Gs. Using entropy as the measure of complexity, this random, almost certainly nonfunctional genome would be considered more complex than the human genome.
In conclusion: the most complex entities are not the most ordered or random ones but somewhere in between.
For further reading, see "Complexity: a guided tour" by M. Mitchell.
Sunday, 8 August 2010
Why humans are different from rats?
Humans have a genome very similar to many other species. For example, more than 90% of our DNA is shared with mice and more than 95% with chimps. Why we are so different from these animals? The evolutionary developmental biology, whose nickname is Evo-Devo, proposes that morphological diversity among species is, for the most part, not due to differences in genes but in genetic switches that are used to turn genes on and off. These switches are the non-coding DNA, or the so-called "junk DNA", which are now known to be used in gene regulation. These genetic networks allow a huge number of possibilities for gene expression patterns, since there are so many possible ways in which proteins can be attached to the switches. The reason the humans share so many genes with quite different species is because, although the genes might be the same, the sequences making up switches have often evolved to be different. Small changes in switches can produce very different patterns of genes turning on and off during the development.
Are you sure this is a gene?
In 2006, a group of 500 scientists were given independently some real DNA sequences and asked whether each sequence qualified as a "gene". For many sequences, the opinion was split: about 60% answered they were confident that the sequences represented genes while 40% were confident that the sequences were not genes. The more specialized a scientist is in molecular biology, the less easy is to define what a gene actually is.
Monday, 2 August 2010
The gene concept and its context
The classical molecular concept of gene is not sufficient to explain several biological processes observed in studies performed in the last two or three decades. The idea of an one-to-one relationship between DNA and protein implies a structural and functional unity; however, the molecular biology showed us that some molecular phenomena like alternative RNA splicing, overlapping genes, and multiple transcription start sites, suggest that the one-to-one relationship is an oversimplification. This concept is no longer useful, except as a handy expression, whose meaning is dependent on the context. Among the several approaches to the multiple usages of gene, one that deserves attention is the concept of the gene as a “fuzzy unity”, which states the genomically diverse nature of the gene. Although this vagueness might have heuristic value, there’s a clear need for more precision. As stated by Eva Neumann-Held in "Cycles of contingency" , whatever the concept of gene is, it should consider the clarification of the purpose and the research context for which the concept is designed.
Wednesday, 28 July 2010
Cancer is influenced by its microenvironment
Recently, there has been an increasing discussion about the role of genes in cancer. The gene-centric view of cancer was reinforced by molecular studies, but this popular conception of the gene as a simple causal agent of cancer is loosing place on the platform of genetic discourse. In fact, cancer is influenced by its microenvironment, yet broader, environmental effects also play a role. Cao et al (Cell 142, 52–64, July 9, 2010) reported that mice living in an enriched housing environment show reduced tumor growth and increased remission. They found this effect in melanoma and colon cancer models, and that it was not caused by physical activity alone. Serum from animals held in an enriched environment (with social and physical activities, joys, etc) inhibited cancer proliferation in vitro. This suggests that the stretch of DNA code for a gene is like a word without a grammar. The environment provides the semantic frame (or grammar) of the organism language.
See here:
See here:
Wednesday, 21 July 2010
Mutational robustness can facilitate adaptation
The relationship between robustness and evolvability is complex because robust populations harbour a large diversity of neutral genotypes that may be important in adaptation. Although neutral mutations do not change an organism’s phenotype, they may nevertheless have epistatic consequences for the phenotypic effects of subsequent mutations. In particular, a neutral mutation can alter an individual’s ‘phenotypic neighbourhood’, that is, the set of distinct phenotypes that the individual can access through a further mutation. Pioneering studies based on RNA folding and network dynamics suggest that genotypes expressing a particular phenotype are often linked by neutral mutations into a large neutral network, and that members of a neutral network differ widely in their phenotypic neighbourhoods. Numerous studies have documented the importance of neutral variation in allowing a
population to access adaptive phenotypes, and neutral networks have consequently been proposed to facilitate adaptation. (Draghi et al. Nature Vol. 463, 21 January 2010).
population to access adaptive phenotypes, and neutral networks have consequently been proposed to facilitate adaptation. (Draghi et al. Nature Vol. 463, 21 January 2010).
Saturday, 17 July 2010
Architecture of complex systems
How is a complex system architectured? To start to answer this though question is necessary firstly to consider that it has large number of relatively simple elements working parallely, which provides robustness and efficiency. Secondly, the system works by simultaneous exploration of many possibilities or pathways, in which the resources are made available according to the probability of success. Many different possibilities are explored simultaneously, but with different depth and speeds. In this process, information is used to evaluate the probability of success and to invest important resources in what is worth exploring.
A good example is provided by Melanie Mitchell in "Complexity: a guided tour". The immune system must determine which regions of the exploratory universe of pathogen shapes will be screened. There are trillions of lymphocytes in the body at any given time, most of them can uniquely identify a specific pathogen shape (or antigen). The shape ranges that are most successful are given more exploration resources: the immunological system increases the number of lymphocytes specifically compromised to that shape range. Thus, the system is able to focus on the most promising pathogens, while never neglecting to explore new possibilities.
A good example is provided by Melanie Mitchell in "Complexity: a guided tour". The immune system must determine which regions of the exploratory universe of pathogen shapes will be screened. There are trillions of lymphocytes in the body at any given time, most of them can uniquely identify a specific pathogen shape (or antigen). The shape ranges that are most successful are given more exploration resources: the immunological system increases the number of lymphocytes specifically compromised to that shape range. Thus, the system is able to focus on the most promising pathogens, while never neglecting to explore new possibilities.
Saturday, 10 July 2010
Cellular automaton
One the most interesting topics in the field of complex systems are the cellular automata, which were invented by John von Newman in the 1940s. The cellular automata (plural of automaton) are grids of cells, where a cell is a simple unit that turns on or off in response to the status of local neighbor cells. There is a rule to update the status of each cell, and this rule is identical to all cells. This rule establishes the status of the cells in the next time step as a function of the current state in its local neighborhood.
At any point in the timeframe, the cellulr automaton processes information by applying its rule to its current configuration. Stephen Wolfram believes that natural systems work much the same way - that they contain information and process that information according to simple rules.
See here a practical example of cellular automata:

Why this idealized model of a complex system is so interesting?
This model simulates complex systems in nature, with no central controller and it can exhibit very complex behavior that is difficult or impossible to predict from the cell update rule.
See here a practical example of cellular automata:

Monday, 5 July 2010
A bioinformatics revolution?
A good question to be answered by those working in bioinformatics is whether there has been a revolution in "techne" ( in the sense of technology) or in "episteme" (scientific knowledge). Or was it in both? I'm affraid Thomas Kuhn could not help us to answer this though question, because maybe the anwer is beyond the limits of epistemology, and the central point is the relationship between science and technology. For those who think about technology as a mere tool for science, it must be said that much of the scientific advancement is due to technology, while technology does not necessarily depend on science to advance. The technology depends on science to discover new phenomena, which will permit the building of new technological tools, but that's not always the case. Technology can advance by combining existing technologies, as usually it does.
In this sense, I like the W. Brian Arthur's definition in his "The nature of technology":
"From all this it follows that science not only uses technology, it builds itself from technology. (...) Science builds itself from the instruments, methods, experiments, and conceptual constructions it uses. Science, after all, is a method: a method for understanding, for probing, for explaining. A method composed of many submethods. Stripped to its core structure, science is a form of technology."
Ad hoc committee
According to Lenny Moss: "The critical decisions made at the nodal points of organismic development and organismic life are not made by a prewritten script program, or master plan but rather are made on the spot by an ad hoc committee".
Molecular biology would be different if its practitioners took this phrase seriously.
Molecular biology would be different if its practitioners took this phrase seriously.
Sunday, 4 July 2010
Putting order in chaos
From "Complexity, a guided tour" by Melanie Mitchell:
"The discovery and understanding of chaos produced a rethinking of many core tenets of science:
1) Seemingly random behavior can emerge from deterministic systems, with no external source of randomness.
2) The behavior of some simple, deterministic systems can be impossible, even in principle, to predict in the long term, due to sensitive dependence on initial conditions."
I think Melanie Mitchell made an useful guide for cancer researchers. Hopefully, they will read her book.
What genes can't do
My second post comes from an article by Richard Levontin, with his broad humanistic perspective of science. It's about the dream of the human genome, and its promises. He wrote:
"Daniel Koshland, the editor of Science, when asked why the Human Genome Project funds should not be given to the homeless, answered, "What these people don't realize is that the homeless are impaired... Indeed, no group will benefit more from the application of human genetics""
Unfortunately, this deterministic ideology is the basis of most genomics research made nowadays. This ontological claim, of the dominance of DNA over all aspects of life, has key social and political consequences.
Hi people!!
This is my first post... nothing to say for a while, but don't worry, there will be a busload of posts pretty soon.
Subscribe to:
Posts (Atom)